
AI
AI for Knowledge Acquisition and Discovery
Developing and evaluating practical human-centered agentic systems and neurosymbolic AI for curation, hypothesis generation, and analysis.
Knowledge-Based AI + Biosystems
Senior computational scientist at Berkeley Lab building knowledge-based AI for complex biological systems.

AI
Developing and evaluating practical human-centered agentic systems and neurosymbolic AI for curation, hypothesis generation, and analysis.

Microbiomes
Building AI-ready, standards-aligned infrastructure for integrating microbiome sequence, metadata, and analysis products across large collaborative ecosystems.
Biological Mechanisms
Representing biological mechanisms in machine-interpretable form to connect genes, pathways, diseases, and phenotypes for reasoning-driven interpretation and hypothesis generation.

Phenomics
Using phenotype-driven inference and cross-species knowledge integration to improve disease interpretation, diagnosis support, and genomic variant prioritization.
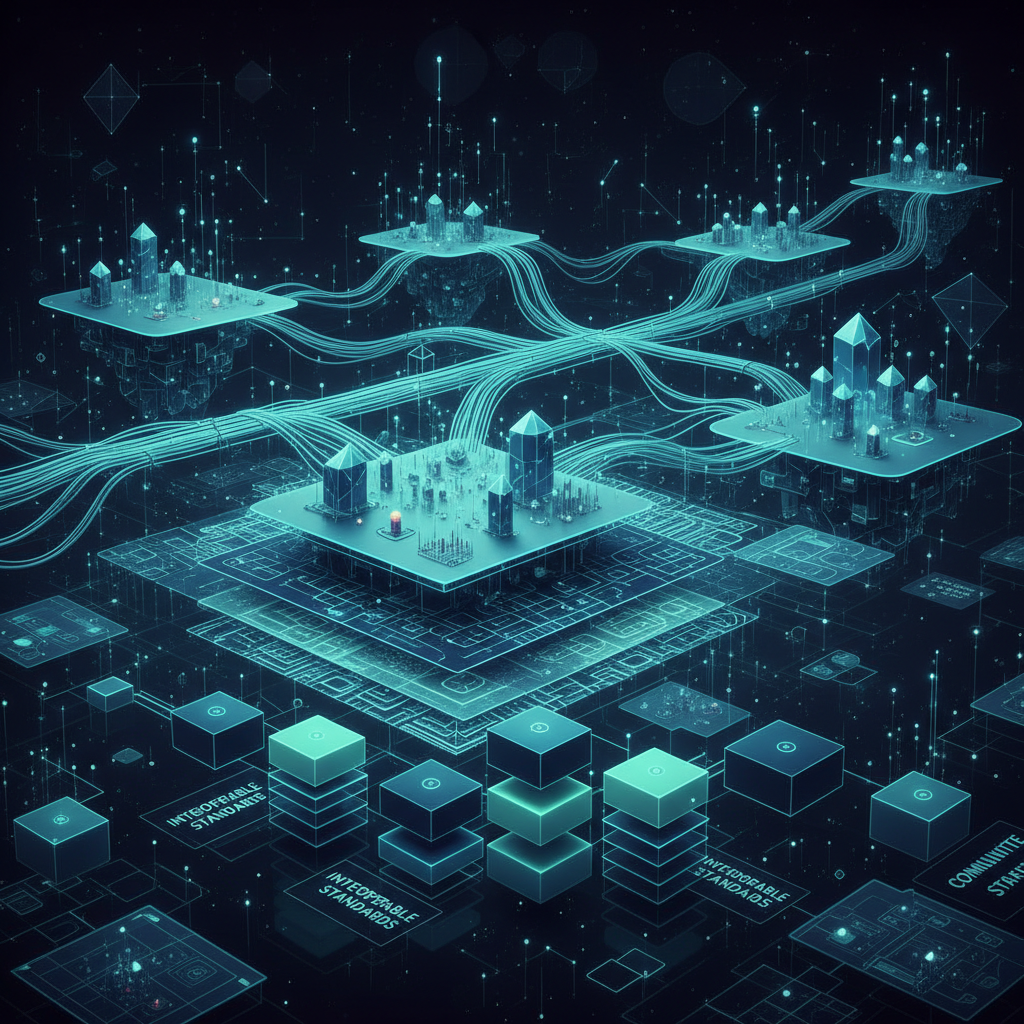

Research Infrastructure
Designing and maintaining standards, ontologies, ontology tooling, and platform resources that make biomedical knowledge and omics data computable, reusable, and AI-ready across communities.

Formal Systems
Developing typed logic frameworks and deterministic reasoning workflows that make biological knowledge explicit, testable, and machine-checkable, and for serving as guardrails for AI systems.
These areas are tightly coupled in practice: standards and ontologies enable robust resources, which in turn support mechanism-aware and phenotype-driven AI workflows.

Mungall CJ, Malik A, Korn DR, ... Hastings J. J Cheminform (2025).
This paper introduces a program-synthesis approach for chemical classification, where LLMs iteratively generate and refine executable classifiers against ontology-grounded examples. It emphasizes transparency and error analysis by exposing classifier logic and failure modes directly in code.
It benchmarks C3PO against SMARTS and deep-learning baselines, showing the tradeoff between maximal predictive performance and interpretable, curator-friendly program outputs that can be audited and improved.

Toro S, Anagnostopoulos AV, Bello SM, ... Mungall CJ. J Biomed Semantics (2024).
DRAGON-AI applies retrieval-augmented generation to ontology authoring workflows, combining ontology context and issue-tracker signals to propose definitions and logical axioms. The study evaluates automated term completion across multiple OBO ontologies with expert scoring and reproducible analysis artifacts.
The focus is practical curation acceleration: ontology editors can use generated candidates as draft artifacts, then accept, revise, or reject with provenance-aware review.

Caufield JH, Hegde H, Emonet V, ... Mungall CJ. Bioinformatics (2024).
SPIRES presents a schema-first extraction pipeline that uses recursive prompting plus ontology grounding to populate structured knowledge bases from unstructured text. It is implemented in OntoGPT and evaluated on nested schema extraction and biomedical relation tasks.
By enforcing schema constraints and grounding to external resources, the method targets higher precision and reproducibility than free-form extraction workflows.

Moxon SAT, Solbrig H, Harris NL, ... Mungall CJ. Gigascience (2025).
This paper describes LinkML as a schema language and tooling framework for creating machine-readable, semantically aligned data models that can generate downstream artifacts such as JSON Schema, OWL, SHACL, and Python classes.
It emphasizes reusable model design and FAIR-aligned interoperability across heterogeneous scientific domains, including biomedicine and environmental data integration.

Joachimiak MP, Caufield JH, Harris NL, ... Mungall CJ. ArXiv [preprint] (2024).
This preprint presents TALISMAN, an LLM-based approach for summarizing gene sets by generating both narrative interpretations and ontology-grounded term lists.
The work evaluates model and prompt variants against standard enrichment baselines, showing where LLM summarization is informative and where precision-recall tradeoffs require careful review.
Beyond these highlights, recurring themes include GO and Monarch resource updates, ontology quality control, and applied agentic AI for biocuration and knowledge extraction.
For more: Google ScholarPubMed / MyNCBIORCIDZenodo Publications
Knowledge Extraction
Schema-constrained extraction tooling for turning biomedical literature into structured knowledge, including SPIRES-style recursive extraction pipelines.
Role: Maintainer and contributor
AI-Assisted Curation
LLM-driven curation assistant workflows for ontology editors and knowledge engineers, focused on draft generation plus human review loops.
Role: Maintainer and contributor
Ontology Toolkit
Unified Python and CLI toolkit for ontology search, graph operations, mapping generation, quality control, and adapter-based access across ontology backends.
Role: Core contributor


Reasoning Engine
OWL EL reasoner designed for incremental and concurrent reasoning, with support for OWL RL and selected SWRL features in ontology-intensive applications.
Role: Contributor
Additional works include ontology QC pipelines, data harmonization tools, and reusable datasets that support curation and analysis across OBO and Translator-adjacent ecosystems.
For more: GitHubMonarch Initiative OrgINCATools OrgZenodo SoftwareTopic: monarchinitiativeTopic: geneontologyTopic: ai4curationTopic: obofoundry
Semantic Resources
Selected ontologies I lead or contribute to.
Browse ontology ecosystem
go-ontology
Core source ontology files for GO, supporting standardized representation of molecular function, biological process, and cellular component for functional genomics workflows.



mondo
Unified disease ontology harmonizing disease concepts across major biomedical resources for robust cross-reference mapping, translational interpretation, and reasoning.
namo
Semantic model and ontology-focused schema for representing new approach methodology concepts in interoperable, machine-readable form.
valuesets
Standardized enumerations and value sets for science and biomedicine, with ontology-linked semantics and LinkML-native generation pathways.
My ontology work emphasizes pattern-driven modeling, review automation, and interoperability governance, including term request triage and coordinated releases across collaborating ontology teams.

Cross-Species KG
The Monarch platform integrates phenotype, disease, and genotype knowledge into queryable graph infrastructure for diagnosis support and discovery.

Functional Knowledgebase
The GO knowledgebase combines ontology structure and high-quality annotations to support enrichment, interpretation, and mechanistic biological analysis.
Disease Mechanisms KB
Curated knowledge base of disease pathophysiology with structured literature-backed claims, phenotype links, and mechanism-centric evidence views.
Queryable Ontology Layer
Standard SQL and SQLite representations of OWL/RDF ontologies that make large ontology knowledge resources directly queryable in relational workflows.
These knowledge resources are designed as interoperable infrastructure, linked by common schemas and standards so data can move across portals, APIs, and computational workflows.
Interoperability
Core standards for representing change, schemas, and mappings across ontology and knowledge graph workflows.
Standards contextChange Representation
Knowledge Graph Change Language (KGCL) is a standard data model and controlled natural language for representing ontology and graph edits as structured change objects.
Schema Standard
LinkML provides a schema language and toolchain for building machine-readable, semantically grounded data models that compile to multiple downstream artifacts.
Chemical Data Standard
Chemical Entity Materials and Reactions Ontological Framework (CHEMROF) defines a LinkML-first schema for chemistry entities, mixtures, and reactions aligned to ontology-driven workflows.
Standards for Genomics and Metagenomes
MIxS (Minimum Information about any (x) Sequence) provides interoperable metadata checklists for genome and metagenome data, improving reusability and cross-study comparison.
Exchange Standard
Knowledge Graph Exchange (KGX) provides a canonical model and toolkit for converting and validating biological knowledge graphs across common graph formats.
Registry Standard
A structured registry for knowledge graphs and related products, with standardized metadata for discovery, interoperability, and lifecycle tracking across resources.

Knowledge Graph Standard
Biolink Model defines shared biomedical classes, predicates, and association patterns to support interoperable knowledge graphs, including NCATS Translator components.
I also contribute to Phenopackets, including work on the schema repository, the core paper, and a LinkML schema view.
Other standards work includes GFF3, AgBioData recommendations, MIRACL (paper), and Mapping Commons. Honorable mention: Bioregistry (paper).
Funding
Selected grants from ORCID and agency records, centered on ontology infrastructure, phenotype interpretation, and AI-ready data ecosystems.
View ORCID record
NHGRI Grant
Supports curation, ontology engineering, and infrastructure sustaining GO as a core, computable functional genomics resource.
Role: Investigator
NHGRI Grant
Builds cross-species phenotype resources and knowledge integration workflows to improve interpretation and prioritization of genomic variants.
Role: Investigator
NIGMS Grant
Sustains BioPortal as a community ontology infrastructure for browsing, search, and machine-accessible ontology services used across biomedical data ecosystems.
Role: Contributor for ontology interoperability and integration
Rare Disease Resource
Phenotype-driven rare disease variant prioritization and case reinterpretation platform integrating HPO-based profiles with variant evidence and cross-species knowledge.
Role: Collaborator and ontology integration contributor
NHGRI Resource Infrastructure
Integrated genome resource platform harmonizing model organism and human data with shared curation pipelines, common schemas, and interoperable knowledge services.
Role: Contributor for ontology and integration standards

NCATS Grant
NCATS Translator-phase funding for translational knowledge graph and ontology infrastructure, aligned with the Common Dialect program line.
Role: Investigator
DOE BER Project
Develops an extensible agentic AI layer over the BER data lakehouse to support harmonized genome-to-phenotype discovery workflows across heterogeneous BER resources.
Role: Project contributor
DOE Office of Science
Community-driven infrastructure for FAIR microbiome data integration and multi-omics interoperability across the DOE microbiome research ecosystem.
Role: Aim 1 lead

DOE BER Initiative
Cross-facility standardized data framework for BER-supported structural biology and imaging resources, designed for multimodal interoperability and AI-ready workflows.
Role: Project contributor

NCATS Grant
NCATS Translator funding line for shared infrastructure and service dialects, forming a closely related phase with subsequent DOGSLED program activities.
Role: Investigator and standards contributor

NHGRI Grant
Coordinates pathway and ontology resources to improve consistency and interoperability for pathway analysis and mechanistic interpretation.
Role: Investigator
NSF Grant
Developed reusable ontology frameworks for plant biology and comparative data integration, reinforcing broader OBO-aligned standards.
Role: Investigator
Many of these awards support long-lived community resources and shared infrastructure, with deliverables spanning ontologies, knowledge graphs, standards, and AI-ready data services.
Speaking
Selected recent talks and keynotes from Zenodo covering agentic AI, ontology workflows, and interoperable bioscience knowledge systems.
Browse talks
Invited Talk - 2026
Practical overview of current agentic AI patterns for biocurators, focusing on sustainable workflows, tooling choices, and near-term adoption strategy.

Workshop Talk - 2025
Foundational session on using coding agents in ontology workflows, with hands-on patterns for high-quality curation and schema-aware editing tasks.

Conference Talk - 2025
Presentation on practical AI-augmented methods for LinkML-based integration pipelines, from schema design to downstream validation and generation.

Featured Keynote - 2025
Keynote on collaborative human-agent workflows for ontology engineering, including design principles for provenance, coordination, and review.

Featured Keynote - 2025
Perspective keynote on combining open knowledge resources with agentic systems while mitigating hallucinations through curated semantic infrastructure.

Featured Keynote - 2025
Keynote focused on practical evaluation criteria for ontology-focused agents and how to benchmark reliability in real curation workflows.

Featured Keynote - 2021
ICBO 2021 keynote on unifying biomedical knowledge graph efforts across ontologies, standards, and community curation pathways.

Featured Keynote - 2020
WOP keynote on cross-ontology design pattern harmonization and strategies for scaling consistent modeling across life science domains.
The talk archive spans keynotes, workshops, and technical deep-dives on ontology engineering, interoperable knowledge resources, and reliable AI-assisted curation workflows.
Learning
Hands-on tutorials and workshop materials, including the ICBO 2025 agentic AI tutorial and GO AI workshop series.
Zenodo presentations
ICBO Tutorial
ICBO 2025 tutorial materials for agent-assisted ontology curation workflows, with practical guidance, exercises, and social coding patterns.

GO AI Workshop
Tutorial material focused on practical AI support for GO curation workflows, including ontology, standards, and social coding pathways.

GO AI Workshop
Foundational workshop on integrating coding agents into ontology development and GitHub-centric review workflows for large semantic projects.

GO AI Workshop
Follow-on hands-on session covering annotation review actions, evidence checks, and quality control patterns for GO standard annotations.
Companion Resources
Operational setup guides and practical checklists that complement the tutorial decks with reproducible workflows and tooling instructions.
Training materials continue to expand across conference tutorials, focused GO sessions, and practical docs designed for day-to-day ontology and curation workflows.
For more: ICBO Tutorial HubZenodo PresentationsAI4Curation Docsai4curation GitHub
Automation
Agent skills and tooling for reliable AI-assisted ontology, schema, and knowledge-base curation workflows.
Browse agentic toolingReference Validation
Validates whether supporting text in structured records is actually present in cited references, helping enforce evidence-backed curation.
Term Validation
Checks LinkML schemas and datasets that depend on external ontologies and controlled terms, improving consistency for agent-generated outputs.
Agent Skills
Reusable skill packs for ontology and biocuration tasks, designed to make agent behavior more consistent, transparent, and domain-aware.
Agent API Bridge
MCP server wrapping GO-CAM editing capabilities, enabling agentic interaction with Noctua/Barista workflows through a standardized interface.
This section tracks practical agentic infrastructure: validators, skills, provenance tools, and MCP wrappers that make AI-assisted curation reproducible and reviewable.
For more: GitHub TopicLinkML GitHubai4curation GitHubAI4Curation Docs
Commentary
Short perspectives on emerging AI for biology, ontology practice, and research infrastructure across social and long-form channels.
View Bluesky perspectives
Bluesky Perspective
Commentary on AlphaGenome and what large foundation models imply for practical biological knowledge workflows, curation, and downstream interpretability.

Bluesky Perspective
Notes on rBio and reasoning-oriented biological AI, emphasizing implications for ontology-aware reinforcement and transparent scientific interpretation.

Interview
Interview and profile discussion covering ontology engineering, biological knowledge graphs, and how AI methods can be grounded in structured semantics.

Bluesky Perspective
Perspective thread on practical training pathways for biocurators, ontology developers, and resource leads adopting agentic AI tools.

Blog
Long-form posts on ontology engineering, curation practice, standards decisions, and the practical realities of semantic infrastructure work.

Article
Practical reflection on autonomous research loops and iterative agent workflows, with concrete takeaways on reliability and operator oversight.

Paper Perspective
Perspective article on automation-ready biology workflows and what robust self-driving lab ecosystems require in standards, data, and semantics.
These perspectives connect day-to-day practice with broader trends in AI, ontology engineering, and scientific infrastructure, from short threads to paper-length commentary.
For more: BlueskyKnowledge Graph InsightsWordPressDEVPubMedORCID